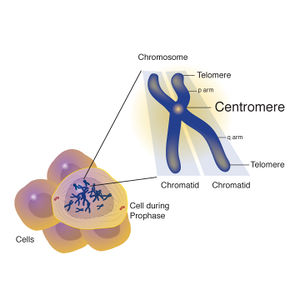
Centromere
From ISOGG Wiki
A centromere is a constricted region of a chromosome that separates it into a short arm (p) and a long arm (q). During cell division, the chromosomes first replicate so that each daughter cell receives a complete set of chromosomes. Following DNA replication, the chromosome consists of two identical structures called sister chromatids, which are joined at the centromere.
| This image is taken from the Talking Glossary of Genetic Terms and is reproduced courtesy of the National Human Genome Research Institute. |
Contents
Centromere locations (Build 37)
The information in this table was extracted by Kitty Cooper from: http://genome.ucsc.edu/cgi-bin/hgTables Instructions were taken from: http://www.biostars.org/p/2349/. Note that the positions in Kitty's chart differ from those provided by Family Tree DNA which we are informed currently uses Build 36. See the FTDNA FAQ Where are the centromeres located on each autosomal DNA chromosome.
| Chromosome | Start | Stop |
|---|---|---|
| 1 | 121535434 | 124535434 |
| 2 | 92326171 | 95326171 |
| 3 | 90504854 | 93504854 |
| 4 | 49660117 | 52660117 |
| 5 | 46405641 | 49405641 |
| 6 | 58830166 | 61830166 |
| 7 | 58054331 | 61054331 |
| 8 | 43838887 | 46838887 |
| 9 | 47367679 | 50367679 |
| 10 | 39254935 | 42254935 |
| 11 | 51644205 | 54644205 |
| 12 | 34856694 | 37856694 |
| 13 | 16000000 | 19000000 |
| 14 | 16000000 | 19000000 |
| 15 | 17000000 | 20000000 |
| 16 | 35335801 | 38335801 |
| 17 | 22263006 | 25263006 |
| 18 | 15460898 | 18460898 |
| 19 | 24681782 | 27681782 |
| 20 | 26369569 | 29369569 |
| 21 | 11288129 | 14288129 |
| 22 | 13000000 | 16000000 |
| X | 58632012 | 61632012 |
Reporting segments crossing the centromere
FamilyTreeDNA allow for a segment crossing the centromere to count as a single segment rather than two separate segments.[1]
Further reading
- Dark centers of chromosomes reveal ancient DNA. Science Daily, 18 June 2019.
- The Y-chromosome's uncharted regions by Sarah Zhang, The Atlantic, 21 March 2018. How nanopore sequencing is making it possible to sequence the repetitive regions in the centromere of the human Y-chromosome.
- GRCh38: Incorporating Modeled Centromere Sequence GenomeRef blog, 14 January 2014.
Scientific papers
- Miga1 KH, Newton Y, Jain M et al. Centromere reference models for human chromosomes X and Y satellite arrays. Genome Research 2014. Published online 5 February.
References
- ↑ Canada R. Message posted in the Family Tree DNA Forums, 27 March 2014. As of 8 January 2020 the out-of-date FAQ. How does the Family Finder program account for the centromere? in the FTDNA Learning Center has not been updated.
|
This article uses material in the public domain from the Talking Glossary of Genetic Terms and is reproduced courtesy of the National Human Genome Research Institute. |