Y-DNA project help
From ISOGG Wiki
Newcomers of Y chromosome DNA tests and in Y-DNA projects (Haplogroups, Surname, Geographical) are often overwhelmed by the many technical terms used for testing Y-DNA listed here and in the Genetics Glossary.
Contents
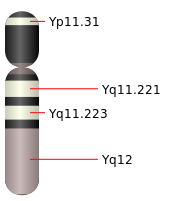
Y chromosome
The Y chromosome (Y-DNA) is a DNA structure found in the nucleus of a male cell. Humans have 23 pairs of Chromosomes, 22 pair of autosomes and one pair of sex chromosomes, XX for females and XY for males. The Y chromosome is passed on without recombination by a father to his sons.
SNP / Variant
A SNP (Single-nucleotide polymorphism) happens when a single place in the genome sequence is altered during the cell formation process and this mutation persists in the progeny (autosomally at least in 1% of the population). A person has many inherited SNPs that together create a unique DNA pattern for that individual. SNPs (Snips) for uniparental inheritance (Y, mt) clarify the branching of different subhaplogroups and allow to discover deep uniparental ancestry. A terminal SNP is the defining SNP of the latest subclade known by current research. It should be unique (UEP) and constant in time. ISOGG mantains a Y-SNP Index where synonymous names are listed. A SNV (single-nucleotide variant) or Variant is a variation from the reference genome in a single place without any limitations of frequency and so might be the more correct term for seldom mutations defining terminal haplogroups.
Clade
Clade comes from the Greek word Klados = branch. A Clade on the Y chromosome tree is also called a Haplogroup. Subclade describes a sub-clade being downstream (occurring later in time). A Clade includes all the descendants of a single MRCA (most recent common ancestor). See also TMRCA.
Upstream, Downstream
In the Y-tree older nodes (ancestors, toward the root) are Upstream. Younger nodes (descendants, toward the present) are Downstream.
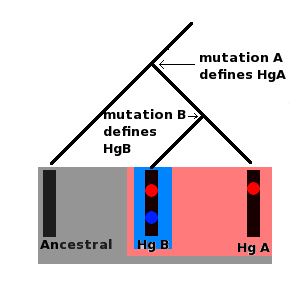
Ancestral, Derived
Ancestral state means a sample is not positive for a certain SNP/Variant (so in that place like the reference sequence). Derived means a sample is positive for a certain SNP/Variant.
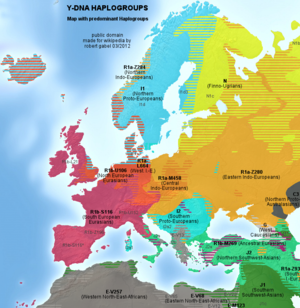
Haplogroup
A Haplogroup is a branch of the human family tree. All men in the same Y-DNA haplogroup share the same SNP or SNPs (unique marker/s in the Y-chromosome) which they have inherited from their common ancestor. The haplogroup is like a name for that common ancestor person. The haplogroup allows to research the modern distribution and the construction of a migration hypothesis of the descendants of the haplogroup founder. The major Eurasian Y-DNA-haplogroups (E1b, G2a, I1, I2, J1, J2, N, O, R1a, R1b, etc.) formed over tens of thousands of years, typical African Y-haplogroups like A00, A0, A and B have even deeper roots. Since 2012 more and more recent SNPs (under 3,000 years old) are discovered and available for research. These types of SNPs are informative for the historical time and allow also research in to the genealogical time.
- Equivalent SNPs: mutations observed in the same haplogroup are equivalent and can all be used to describe a haplogroup. It is impossible to define the chronological order (time of occurrence) of the SNPs in one haplogroup. Example: L21/M529/S145, L459, FGC3218/S552/Y2598;
- Synonymous SNP: names describing the same mutation are synonymous; example: L21 = M529 = S145 often listed as L21/M529/S145;
Nomenclature System (YCC)
2002 the Y Chromosome Consortium (YCC) proposed two widely accepted nomenclature systems for Y-DNA haplogroups.[1] Major haplogroups are labeled with large capital letters (A–T). Examples here for the haplogroup defined by the SNPs L21/M529/S145 and L459:
- Hierarchical system: R1b1a2a1a2c (ISOGG 2016 11.20), R1b1a2a1a1b4 (FTDNA 2009), R1b1a2a1a1b3 (ISOGG 2012 v7.62), R1b1b2a1a2f (23andMe 2009).
- Shorthand - SNP system: R-L21, R-M529, R-S145. This system is more robust to changes in topology but widespread SNPs have often up to three synonymous names. Additionally different corporations/labs in many cases select an equivalent SNP for the same haplogroup as primary/defining (R-L459). For seldom and new terminal SNPs there is also the risk that they are not unique (recurrent, unstable) or not detectable with all lab methods.
- Basic Hierarchy + Shorthand system: since 2013 this system is used by some publications to show the basic hierarchy under a main haplogroup combined with a SNP of a subclade deeper down then the listed hierarchy: R1b-L21, R1a-L664. Especially for unknown SNP names this allows easier recogniation of the basal position. Stable basal Haplogroups names are limited and might be: E1b, G2a, I1, I2a, I2b, J1, J2a, J2b, R1a, R1b.
- Paragroups are distinguished from haplogroups by using the * (star) symbol, which represents chromosomes belonging to a clade but not its researched subclades defined in the same publication: R-L21*. When a paragroup is mentioned outside an accompanying publication it is better to mention the excluded subclade/s by SNP name in parenthesis after an x: R-L21(xDF13,DF63)
Name Versions - Y-Chromosome Phylogenetic Trees
Since 2002 many new ramifications (SNPs) even in basal branches and subclades where found. The YCC [2], other scientific papers, societies and companies published substantial refining and updates to the Y-Chromosome Phylogenetic Tree, where the haplogroup names for deep clades often are different. In Genetic Genealogy the following name versions are important:
- Since autumn 2012: many scholars, companies and genetic genealogists agree [3] that the Shorthand - SNP system is the solution to avoid confusion for the future. FTDNA totally merged to this system too.
- FamilyTreeDNA/FTDNA:
- from 2005 to 2012 uses mainly the Hierarchical system; in some places the Shorthand - SNP system. 2005 Y-Tree PDF, 2008 Y-Tree PDF
- 2009-2014 in myFTDNA and FTDNA Projects a slightly updated YCC 2008 version was used, ytree.ftdna.com (including Draft version, now offline)
- from April 2014 to 2018 FTDNA uses a Phylogeny created and based also on Genographic Project Y-SNP results. The tree has some bugs (mainly with recurrent terminal SNPs) and useful ISOGG SNPs and phylogeny known before 2014 are omitted.
- in September 2018 FTDNA released the public Y-DNA Haplotree with listing of amount of postive samples and countries for haplogroups; in January 2019 FTDNA for BigY customers released the Big Y Block Tree (not public) a vertical-block diagram which is based on Big Y results with detailed SNP results.
- YFull public YTree: v3.07 from 28 March 2015 and since then updated almost monthly. Based on Next Generation Sequencing samples from scientific studies (if enough coverage) and private DTC testing (for a fee). Links to major Haplogroups: E1b-P177, G2a-P15, I1, I2, J2a-M410, J2b-M102, LT, R1a, R1b-S250/DF27, R1b-S145/M529/L21, R1b-U152/S28, R1b-U106/S21/M405
- ISOGG: the Y-DNA Haplogroup Tree since 2006 is updated according to new evidence from publications and public research (Y-DNA Projects etc.). Unfortunately the manual updating was increasingly unable to monitor the huge amount of updates at least for some Haplogroups since at least 2014 when Next Generation Sequencing became widespread for the Y-DNA. Most used reference: E, G, I, J, R.
- Citizen science research trees: E1b (Forum with Links), G (login needed), J1 (2014), J2 (mainly 2014-2017), R1b-U106/S21/M405 (Yahoo Group offline?), The Big Tree R1b-P312/S116 (Alex Williamson, since 2019 less activity)
- Phylotree/Y: minimal reference phylogeny for the human Y chromosome (population studies, forensic labs, etc.)
- 23andMe: in use since 2009. No public version is available. No major updates so very basic. If you are a customer you can view the linked information: Paternal Haplogroup Tree, Haplogroup Tree Mutation Mapper
Y-STR - DYS values
STR (Short tandem repeat) is a short DNA motif (pattern). Y-STRs occur on the Y-DNA. DYS (DNA Y-Chromosome Segment) numbers show the repeats of an STR on that position. A DYS value tipically mutates with a certain (low) probability to a higher or lower value from generation to generation. By this DYS values are not unique and not constant in time.
Y-DNA - Haplotype
Y-DNA Haplotype is defined as one person's set of values for the DYS locations. A set of DYS values is highly informative for tracing recent ancestry (genealogical time). The quantity of needed DYS values depends on the research goal and the frequency of nearby haplotypes. For surname projects 12 or 25 markers can be enough, while for extended haplotpye studies (lineage distinction, pre surname time, SNP research) and to find more distant matches 37 to 111 markers are used. Modal haplotype is the most commonly occurring haplotype derived from a specific group. It should be near or identical to the Haplotype of the MRCA (most recent common ancestor) of that group.
Y-DNA - Matches
Y-DNA Matches are other kits (tested males) that have the same or similar numbers for the DYS values. While the same values usually are only probable in near relatives (father, son, brother, grandfather, cousins), step mutations can show a relation until many generations ago. In major european haplogroups (R1b-U106, R1b-U152, R1b-L21, I1-M255, E1b-M78, J2a-L26, G2a-L30, I2-M223, etc.) many subclades have overlapping haplotypes. In this cases only by high DYS coverage and positive testing of a recent terminal SNP the recent common ancestor is proven. See also TMRCA.
Cluster
A group of kits all together near by haplotype (Y-DNA matches) and having no discovered unique SNP defining a haplogroup is called Cluster. The given name is only temporary defined and usually only used within the project. It is interesting for members of a Cluster to work together for discovery of new SNPs (WTY, Y-Sequencing, Deep Clade test) and test funding of interesting and informative low coverage samples.
Y-STR testing
See Y-DNA STR testing chart, Y chromosome DNA tests
FTDNA Y-STR DYS Upgrade (Y-Refine)
- Login to FTDNA (MyFTDNA) > Order an Upgrade > Order a Standard Test > Select A Product
- Select your desired Upgrade:
- 12 Marker kits: Y-Refine12to25, Y-Refine12to37, Y-Refine12to67;
- 25 Marker kits: Y-Refine25to37, Y-Refine25to67;
- 37 Marker kits: Y-Refine37to67, Y-Refine37to111;
- 67 Marker kits: Y-Refine67to111
- Make your order clicking on Next, etc.
Y-SNP testing
See Y-DNA SNP testing chart, SNP testing
If a terminal SNP can be predicted with high probability or not is depending on the haplotype frequency and number of DYS values. For a strong prediction, usually possible for seldom haplotypes with enough matches and kits with 67 or 111 markers, the testing of single SNPs is often the better choice. If a kit has low DYS coverage (12 to 37 markers), has no SNPs tested and matches to a huge haplotype cluster from different subclades a SNP test covering many SNPs at once is often the smarter solution. SNP chips provide a good value for the money, while Y-sequencing is the method providing deepest results down to "family haplogroups".
FTDNA Single SNP test - Order an Advanced Test
- Login to FTDNA (MyFTDNA) > Upgrade Button > Advanced Tests - Buy Now > Select Filter Test Type SNP Marker
- Search for recommended SNP (for example L123) and Add it to the cart. Repeat the last step for additional SNPs.
- Make your order clicking on Next, etc.
Since 2015 new deep clade SNP testing panels are offered.
YSEQ Single SNP test (Sanger Sequencing)
Thomas and Astrid Krahn since November 2013 over YSEQ offer a menu of single SNPs to order. New SNPs (so far discovered only through SNP chips or Y-sequencing) can be requested. Also SNP panels are available.
SNP chips: Ancestry.com, 23andMe, MyHeritage
The tests are optimized for low costs and comparison regarding near relatives and genetic population ancestry by using auDNA and X-DNA, see Autosomal DNA testing comparison chart. Depending on the Chip Version the results contain only a limited selection of Y-SNPs and also data for the mtDNA. By using tools a basic Y-haplogroup level can be predicted, which is usually not usable for genetic genealogy. See Y-SNP_haplogroup_prediction_tools (YSEQ Clade finder , Chris Morley's Y-SNP subclade predictor, etc.)
Historic / Vendors in the past
Website until late June 2020: Genographic Project: Geno 2.0 SNP chip Geno 2.0 test is available at the Genographic Project page and tests for over thousands of known Y-SNPs. It includes SNP results for mtDNA, auDNA and X-DNA. Website after June 30, 2020 probably offline. If you have received your results please transfer your data from Genographic Project Geno 2 (in Profile / Expert Options) for free to Family Tree DNA. Only by doing this Y-SNP results will be available in FTDNA projects. Download also your raw data for possible future use.
Until 3 July 2017: BritainsDNA Chromo 2 Raw Y-DNA SNP chip BritainsDNA and its associated companies (ScotlandsDNA, IrelandsDNA, YorkshiresDNA and CymruDNAWales) looked at thousands of Y-SNPs. This test was assumed to bring good results for haplogroups common in British Isles and Netherlands. The user database was closed 31 August 2018. Note that BritainsDNA and the companies under its ownership ceased operations and the See the ISOGG Wiki entry for BritainsDNA to learn more.
FTDNA Settings: Paternal Ancestor Info
For every Y-DNA project it is very informative and sometimes important if the oldest paternal line (biological father of the father of the father etc.) information is given completely. This person is also called most distant known ancestor in the paternal line (Y-MDKA). This setting can be entered/changed in myFTDNA > My Account > Most Distant Ancestors: Direct Paternal:
- Country of Origin: enter the oldest known or strongly suspected country of origin of your Y-DNA. For example: "Germany". Otherwise enter "Unknown Origin".
- Name: enter name, year born/dead, municipality/city/town/village, province/county/region; example: "John Schmid, 1788-1852, Augsburg, Swabia". If only one date is known add informations like "b. 1788", "d. 1852", the place information should be the oldest known (usually birth, then marriage or other records and then dead). If there was a known Non-paternity event (surname change) and the biological father is unknown please indicate the latest known male ancestor and his mother ("Martin son of Maria Miller, b. 1822, Augsburg, Swabia"). It is a common error that the paternal ancestor of Maria is indicated ("Georg Miller, 1730, Swabia") because this ancestor is not the genealogical Y-DNA ancestor (strict paternal line).
- Ancestral Location Direct Paternal: enter the full location information: house/place/street, municipality/city, ZIP, province/county/region, country. For example: "Hauptmarktplatz, Hoher Weg, Augsburg, Swabia, Bavaria, Germany". Enter the coordinates: Latitude and Longitude
GEDCOM/Family Tree
For surname projects or when your paternal ancestors did often change location (region, countries) the uploading of a GEDCOM file (see Genealogy software) with all the paternal ancestors included will greatly help research and the finding of new connections. You help especially adoptees or unknown male offspring to find the possible generation of connection.
Privacy & Sharing
To allow public viewing and sharing of Y-DNA results since 2015 for new kits it is necessary to change the new presetting "Project Members" under Privacy Settings > My DNA Results - Select who can view your DNA results:
- Who can view my DNA results in group projects? change to Anyone
This is a very important setting to help Admins of Surname DNA projects and Geographical DNA projects to get the best advise of terminal grouping from Y-DNA haplogroup projects.
Y-Sequencing
A Next Generation Sequencing (NGS) readout of the Y-DNA and analysis of the data is the best available option to get all virtually available information on the Y-chromosome. If the coverage is good enough, all known SNPs can be checked and additionally new SNPs will be found. Since 2013 this service is available DTC (Direct To Consumer). For comparison see Y-DNA SNP testing chart, Y-DNA next generation sequencing.
Family Tree DNA Big Y
Y-DNA Sequencing is offered by Family Tree DNA. After test results are complete, some raw data (VCF, BED) is made available with a download link which can be found on the "Other Results" > Big Y Results page. Trough a form request the BAM file including all useful sequence data will be made available. See Y-sequence. These file can be productively examined or analyzed by those with the necessary expertise, such as certain project administrators, FGC (Full Genomes Corporation), and YFull. See below for more about FGC and YFull.
FGC Comprehensive Y-Chromosome Sequencing
Y-DNA Sequencing including SNP and STR Reports as well as a FMS is offered by Full Genomes Corporation. Different tests including the Y are available: Y-Elite as most comprehensive DTC test and various whole genome tests. As noted above, FGC also offers Interpretation of BAM Files. Because of the GDPR services are not available in the EU.
YSEQ Whole Genome Sequences
YSEQ has introduced various NGS Tests which are optimized for genetic genealogy especially regarding the Y-DNA. Comprehensive Interpretation of the raw data is provided to facilitate the usage and comparison.
WGS: Dante Labs, Nebula, etc.
Entry level prices for Whole Genome Sequences where made available by various providers: Dante Labs, Nebula Genomics, Veritas Genetics, see List of DNA testing companies. Long pending results and low customer care are reported often. Combined with the fact that genetic genealogy Interpretation is not provided, these tests are only suggested for experts or if one is giving support.
YFull interpretation Y-Chromosome sequence
YFull offers analysis of Y-DNA sequences (raw data/BAM files) for $49 (public samples for research and interpretation). Technical requirements raw files: Alignment BAM file, Coverage min 30X, Read length min 100 bp.
- Haplogroup, SNPs, YTree
- STR results: All known (over 440) Short Tandem Repeats extracted from Y-Chromosome (over 100 new)
- Private/Novel SNP results: SNPs found only in your sample (comparison is done automatically with all other YFull samples)
- and other services
References
- ↑ Y Chromosome Consortium (2002-02). "A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups". Genome Research. doi:10.1101/gr.217602. http://genome.cshlp.org/content/12/2/339.full. Retrieved 2012-04-12.
- ↑ Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (2008-05). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. doi:10.1101/gr.7172008. http://genome.cshlp.org/content/18/5/830. Retrieved 2012-04-12.
- ↑ Let's All Start Using Terminal SNP Labels Instead of Y Haplogroup Subclade Names, Okay? http://www.yourgeneticgenealogist.com/2012/09/lets-all-start-using-terminal-snp.html
See also
- Y-DNA haplogroup projects
- Y-DNA SNP testing and haplogroups FTDNA Learning Center