DNA in health and disease
From ISOGG Wiki
DNA testing is a great new technology! Learn about yourself and your health! Your life will be better! So goes the pro-argument – typically from those who want to sell you a DNA test. There are also those who are sceptical. Can you really get anything out of it, that DNA test? Is there any reasonable benefit that a healthy adult could ever hope to get from DNA testing?
That is the subject I would like to write about, because the answers to those questions actually have quite a lot of nuances. If you've just bought an ancestry test and are interested in checking out the health aspects, these are the questions you'll want to ask.
The first challenge in understanding is that it is very rare for DNA to contain anything that has a 100% chance of anything. It's self-evident when you think about it: How could anyone – or indeed any organism – ever manage to survive in a chaotic world if the path of your entire life trajectory was pre-determined from the beginning? We are more robust and flexible than that, humans, as well as animals, plants and bacteria. DNA is, rather, like an architectural plan providing smart solutions for many different outcomes: DNA describes how a flexible self-learning immune system is created. It describes how to make muscles and fat stores that can grow and shrink according to circumstances. And it's the plan for how a curious and adaptive brain is created and developed to make wise decisions. But DNA itself does not make the decisions for you. And there is absolutely no place where it will ever say that you are going to develop cancer at exactly 36 years of age.
Sometimes, however, there are less optimal variants in our DNA. We call them gene errors, pathogenic variants or simply mutations, and in rare cases these can result in serious hereditary diseases. The reason why these cases are rare is because that is what natural selection is all about: The more serious a gene error is, the harder it is to grow up with it and pass it on to the next generations. That's just how it is.
But then what about the DNA variations that cause small changes in the muscles of your heart, thereby giving you a slightly increased risk of getting a stroke? Or a 2 mm wider waistline? Such small changes have probably not played any major role in the great game of evolution. So, not surprisingly, there are millions of such small DNA changes between people. We have mapped them today and given them names: we call them SNPs. Some of these are the same SNPs that are used by the testing companies to estimate your ancestry percentages or to predict your relationship with your cousins. One of the exciting developments in health genetics these days is that we are learning how to handle the sum of these many small SNPs. We can summarise the combined effect of all these different SNPs to calculate what is known as a polygenic risk score.[1] In almost all cases, the answer is that there is some effect, but that everything is not pre-destined from birth.
Contents
DNA, environment and their interaction
There are people who have variants so bad that they almost surely will get a genetic disease. These diseases all have exotic-sounding names: familial hypercholesterolemia, phenylketonuria, Niemann Pick, Tay Sachs, Ehlers-Danlos Bloom-Torre-Machacek and many more. They are rare and typically discovered at young age. They are the exceptions. Evolutionary biologists can say a lot about why they are exceptions. But they are.
Then there are the more common diseases that most people know the name of: stroke, hypertension, cancer, diabetes, rheumatoid arthritis, schizophrenia and depression. These diseases are unfortunately more common and, yes – you may well be born with a really unfortunate genetic risk of getting them. But this genetic risk is of a different kind than that of the more rare genetic diseases.
This is the second main challenge in genetic health interpretation: understanding the huge difference in impact that different genetic variation can have. Because your DNA determines who you are and how your health develops in interaction with the things you do and are exposed to throughout life. This is your 'environment', as we geneticists like to call it, or just “life”.
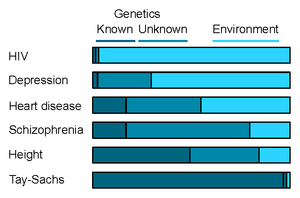
In the figure the heritability of various diseases (and height) is shown. The more dark blue there is in a block, the greater the influence genetics has on that trait. The 'unknown' factor in the slightly lighter dark blue is also genetic, we just haven't yet figured out how it is connected there.
Genetics and environment exist in tandem. That is the case for the vast majority of traits; both in common diseases but also how we are, think and look. When the otherwise super-fit marathon runner gets a stroke as a 45-year-old, or when the chain smoker celebrates their 105th birthday, it is probably the genetics that pulled in the opposite direction.
Very rare genetic variants can have a big effect on you. Commonly found genetic variants tend to have small effects and typically only when considered as a whole in combination with many different SNPs. This is a key point to understand – and is indeed where the genetics of trait and disease prediction is today. There’s a great figure on wikipedia to illustrate this. Go look at it – smart people wrote that Wikipedia article. In the figure, the upper-left area is the rare strong effect genetics. I’ll call them the "rare baddies", because highly penetrant Mendelian mutations will derail all further understanding. The lower-right area represents the small common effects. I’ll call them the polygenic effect SNPs, because they only make sense when summarized over many of them. There are many (poly) SNPs (genic) that are responsible for the effect. Polygenic risk scores were all the rage in genetics research communities in 2018.[3]
The rare baddies
If you really want to go into health and disease interpretation yourself, you need to have these two main categories very clear in your mind. It’s very common to read online about someone who has found that they carry a polygenic effect SNP and then get scared as if they had a rare baddie. They usually didn’t – simply because rare is rare. The options you have on the path of self-analytics therefore flows from these two categories. First the rare category.
If you look at one SNP at a time, you have in a sense already chosen to look for rare genetic disease – rare baddies. Or at least you should think like that, because of the above considerations. There are many online tools available: Promethease, livewello, codegen.eu and xcode.life are a few of the popular ones. Do not use these for any insight into common diseases. If you have heard about the disease before, it’s too common. It will not be affected by a single SNP. Not to any appreciable degree. That’s why the exotic sounding diseases from the previous section are better examples of use-cases. I’m assuming you hadn't heard of those before. The list of them is pretty long and they are all very rare. Promethease, for example, has a very extensive list of SNPs (SNPedia) classified by something they call magnitude. Their magnitude metric (roughly) corresponds to the Y-axis effect size in the Wikipedia map we talked about before. Do use the magnitude recommendations, meaning; do ignore anything with a low magnitude score completely. “Probably worth your time” is an overstatement, but the larger magnitudes are more important.
A key paradox in this analysis mode is that the rarer a SNP is, the less likely it is to be detected by the microarray technology that we use in genetic genealogy. It requires the more expensive DNA sequencing technology, which is not (really, yet) part of consumer genetics and so outside of the scope of this article. Another important consequence is that it becomes impossible to use consumer genetics to exclude the existence of strong pathogenic variants. For any given known variant that destroys a gene, causes diseases and is also measured on a microarray, there will typically be a thousand other possible rare variants in that gene that would cause the same disease but are not measured at all.
If you find anything in this category, then it is crucial to be aware of the accuracy of that finding. Because you are basing conclusions on a single SNP it is obviously important that your information is correct. Two key questions are therefore worth asking. How accurate is the measurement of the SNP? And how accurate is the interpretation of its effect on disease? The measurement precision of any SNP on a microarray is approximately 99.5%.[4] This sounds quite good, and it is – but keep in mind that in your data, with almost a million SNPs reported, it will mean that around 5000 SNPs are wrong. Secondly, the question of interpretation is important. Assuming your genotype is correctly measured, it is also important to find a trustworthy source for how to interpret it. In a study from 2018, researchers from Ambry Genetics found that they disagreed with the medical interpretation of as many as 40% of the pathogenic SNPs highlighted in other third-party analysis software.[5] This underscores that there is still a lot of subjectivity involved even in the interpretation of known SNPs.
The polygenic effect SNPs
Down the other road – the one with the polygenic effect SNPs – there are fewer online analytics options and the interpretations become more difficult. In March 2019, 23andme released a new score of this type for diabetes. This was the first example of a polygenic risk score that was approved by the FDA. But even before this approval some polygenic risk scores were available at third-party websites such as GenePlaza, 24genetics and DNA.land. However, the website with the broadest selection of polygenic risk scoring, is probably impute.me, which is something I have worked a lot with myself – so a disclaimer – I too am a part of this field. I do it because I am very interested in contributing to an increased understanding of how we can use genetic analysis in common disease, which is the subject of this article. No matter how you twist it, genetics will not explain everything about the common diseases that are affected by the polygenic effect SNPs. Not now, not ever. Because environmental factors play a big part in these diseases too. That was the entire point of the figure with the blue bars.
Therefore, good examples of the effective use of polygenic risk scores are few and far between but there have been a handful of published studies.[3][6][7] An analysis may be helpful if you have sought medical advice and there is doubt about a specific diagnosis. If you are being evaluated for several possible diagnoses, it may be helpful to know if you are at a particularly high genetic risk for one particular disorder. This is an area of active research (e.g. this analysis module), and it is not part of routine care. It is, however, part of a complete view of possibilities in the use of consumer genetics in health. And it is without doubt the only correct way to analyse diseases that we know for sure are results of multitudes of different genetic and environmental effects.
Overall, the key point of having a separate section on polygenic effect SNPs is to underscore the difference between how rare genetic diseases with strong single-SNP effects should be analyzed (previous section), and contrast it with how common diseases should be analyzed. The latter types of disease are without exceptions the product of a complex interplay of environment and thousands of different of SNPs. Now and in the future, huge studies will report how we now have new knowledge of thousands more SNPs, potentially all the way up until the unknown genetics part of the blue bar figure has disappeared. But a single SNP associated with a common disease will never have any appreciable effect on your life, not unless it is considered together with the many other SNPs that affect the same disease.
If you make any observations in the polygenic trait category, then it is crucial to be aware of how large a proportion it explains of your overall risk. If we only know few of the SNPs involved in a very complex disease, say depression, then the impact on our life is still only modest.[8] If we know a lot about the genetics of a disease, but it is not very heritable, say HIV, then the impact is also modest. Conversely, if a trait has a high heritability and we know a lot about the SNPs involved in it, say height, then a polygenic risk score can potentially have a sizeable impact on your life.[9] Particularly if you find yourself in the extreme ends of scores.[7] Each of these cases are visualized in the blue-bar figure above. So in summary; if the darkest block is large then you can say something about the genetics of that disease. If it is not, then you cannot.
The intersection between rare and common
Then of course there are the exceptions to these two broad categories of analysis. There always are. A few common diseases have individual SNPs with effects so strong that they impact you more directly. These are breast cancer (BRCA), Alzheimer’s (APOE) and Parkinson’s (LRRK2 and GBA). 23andMe has received approval from the FDA to include reports on these three diseases in their 23andme health product and often highlight them when called to defend clinical utility.[10] This is not by coincidence. In fact, this group of three diseases could be labelled as “rare baddies that are rather common". This is also the reason why APOE is highlighted in the wikipedia figure, where an effect size of x5 is given. Breast cancer variants have a similar effect size, where a damaging variant in one of the BRCA genes will increase the life time risk from 12% to more than 50%.[11] So highly appreciable increases in risk, for diseases that unfortunately are already common. Because these diseases are common and the effect is strong, they are also the ones that are very often discussed. None of the three, however, will result in certain disease (e.g. 50% risk for breast cancer). This is one main point here at the intersection between common and rare genetics. It is still not deterministic. A second main point is that for any other SNP-disease association that you are investigating, you can assume that it is either (a) rarer than these three examples, while still having a strong effect size (the "rare baddie" category) or (b) more common than these three examples, but with a weaker effect on your life (the "polygenic effect SNPs" category).
Another conceptual area that could be called an exception from the rare versus common categories is that of genetic precision medicine. All people respond differently to medical treatments. It would often be good to know a person's drug response up front before starting treatment. It saves on time, money and harmful side effects. For many diseases there is more than one treatment option available, so picking the treatment that is most likely to work for you personally seems smart.[6][12] The use of genetics in optimal drug choice is often termed precision medicine. The reason that this deserves its own section, is that it somewhat changes the risk/benefit calculations of everything we have discussed before. As with common diseases, we usually only know of weak predictive effects for drug responses. But that is a smaller worry than if you were to assign yourself a risk of disease – because the harm of a wrong choice is not so bad. Your doctor simply needs to prescribe a different medication. It is likely that such drug response predictions will be more useful in the future.
What to do with your findings
What happens if you find something in your results? You have come to a conclusion about your health, and you want to do something about it. What then? If it is that important, you must seek medical advice. Consumer genetics does not provide a clinical diagnosis. This advice has been repeated again and again by authorities and yet you are reading this article, so I’m assuming you’ve heard it before too.[13][14] My belief is that since people are going to pursue these options anyway, it is much more useful to provide complete, useful and correct information about the concepts and the state of the art about what we know. Unsurprisingly, I can highly recommend my own book about this. [2]Again a disclaimer – I write books about this. Information is important.
I know many medical doctors who become outright annoyed when presented with consumer genetics results. They should not always be. A suspected disease-causing BRCA variant does need to be followed up. It’s a “rare baddie”. But it has to be interpreted in the right context. I too would be annoyed if I was approached by someone saying that their mental health problems were due to 13 bad genes, of 121 for schizophrenia. These are polygenic effect SNPs, it simply doesn’t make sense to look at them that way.[15] Not unless you know if the effects are cancelling out in your favour or not. I hope this article has given you some insights into these concepts and the context where they can be interpreted, and I hope this can help you on your way, if you choose to go for self-analytics. Despite the caveats.
Finally, please don't make either me or ISOGG responsible for your analysis; this article is merely intended to provide helpful advice in a rapidly advancing field. So – final disclaimers – several apps and companies are mentioned. None of the mentions are recommendations from me or from ISOGG – but we have to discuss what is out there, so I figured an article without names would be pretty pointless. This is not formal medical advice. Consumer genetics never can be. Go see a doctor or a genetic counsellor if you are in doubt.
Further reading
- Polygenic risk scores and tea-drinking. A walk-through of the caveats of translating polygenic risk scores between nationalities by Graham Coop. The Coop Lab blog, 14 March 2018.
- DNA is not our destiny; it's just a very useful tool by Ewan Birney, The Guardian, 5 April 2018.
- The approach to predictive medicine that is taking genomics research by storm by Matthew Warren, Nature News, 10 October 2018.
- Pharmaco-whaat?! Understanding pharmacogenomic testing and how it could aid your doctor in prescribing medication Watershed DNA, 7 January 2019.
- Alzheimer's disease - key points to know in light of the new 23andMe reports Watershed DNA, 7 May 2017.
- Game of chances: inheritance is a question of probability, not destiny by Adam Rutherford, The Guardian, 21 June 2015.
- Forecasts of genetic fate just got a lot more accurate by Antonio Regalado, MIT technology review, 21 February 2019.
Resources
- Health genetics forum - 23andme promethease impute.me codegen.eu livewello. A Facebook group dedicated to discussion of consumer genetics and health implications.
See also
References
- ↑ Matthew Warren. The approach to predictive medicine that is taking genomics research by storm
- ↑ 2.0 2.1 Folkersen et al (2018) Understand your DNA. World Scientific Publishing
- ↑ 3.0 3.1 Khera et al (2018) Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations
- ↑ Hong et al. (2012) Technical reproducibility of genotyping SNP arrays used in genome-wide association studies
- ↑ Tandy-Connor et al (2018). False-positive results released by direct-to-consumer genetic tests highlight the importance of clinical confirmation testing for appropriate patient care
- ↑ 6.0 6.1 Santoro et al (2018) Polygenic risk score analyses of symptoms and treatment response in an antipsychotic-naive first episode of psychosis cohort
- ↑ 7.0 7.1 Lewis and Hagenaars (2019). Progressing Polygenic Medicine in Psychiatry Through Electronic Health Records.
- ↑ Wray et al (2018) Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression.
- ↑ Wood et al (2014) Defining the role of common variation in the genomic and biological architecture of adult human height.
- ↑ Wojcicki (2019) 23andMe Responds: Empowering Consumers New York Times
- ↑ U.S.Preventive Services Task Force (2015). Risk Assessment, Genetic Counseling, and Genetic Testing for BRCA-Related Cancer in Women: Recommendation Statement
- ↑ Cui et al. (2013). Genome-wide association study and gene expression analysis identifies CD84 as a predictor of response to etanercept therapy in rheumatoid arthritis.
- ↑ American College of Medical Genetics (2016) Direct-to-consumer genetic testing: a revised position statement of the American College of Medical Genetics and Genomics.
- ↑ American College of Obstetricians and Gynecologists (2018) Practice Advisory: Response to FDA’s Authorization of BRCA1 and BRCA2 Gene Mutation Direct-to-Consumer Testing
- ↑ Ripke et al (2014) Biological insights from 108 schizophrenia-associated genetic loci.
