Single-nucleotide polymorphism
From ISOGG Wiki
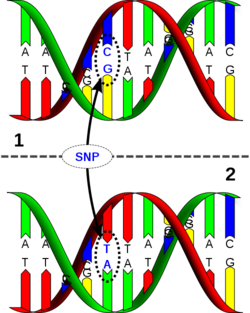
A single-nucleotide polymorphism (SNP, pronounced snip) is a DNA sequence variation occurring when a single nucleotide adenine (A), thymine (T), cytosine (C), or guanine (G]) in the genome (or other shared sequence) differs between members of a species or paired chromosomes in an individual. For example, two sequenced DNA fragments from different individuals, AAGCCTA to AAGCTTA, contain a difference in a single nucleotide. In this case we say that there are two alleles: C and T. Almost all common SNPs have only two alleles.
Within a population, SNPs can be assigned a minor allele frequency — the lowest allele frequency at a locus that is observed in a particular population. This is simply the lesser of the two allele frequencies for single-nucleotide polymorphisms. There are variations between human populations, so a SNP allele that is common in one geographical or ethnic group may be much rarer in another.
Contents
Types of SNPs
Single nucleotides may be changed (substitution), removed (deletions) or added (insertion) to a polynucleotide sequence. Single nucleotide polymorphisms may fall within coding sequences of genes, non-coding regions of genes, or in the intergenic regions between genes. SNPs within a coding sequence will not necessarily change the amino acid sequence of the protein that is produced, due to degeneracy of the genetic code.
A SNP in which both forms lead to the same polypeptide sequence is termed synonymous (sometimes called a silent mutation) — if a different polypeptide sequence is produced they are nonsynonymous. A nonsynonymous change may either be missense or nonsense, where a missense change results in a different amino acid, while a nonsense change results in a premature stop codon. SNPs that are not in protein-coding regions may still have consequences for gene splicing, transcription factor binding, or the sequence of non-coding ribonucleic acid (RNA).
Listing SNPs
SNPs, in particular the key marker SNP used in Hapologroups, are designated by color or code in genetic genealogy.
A non-tested or estimated SNP is usually designated in a black or the standard color of the text or without a symbol of tested confirmation ( + ) or non-confirmation ( - ) code.
A tested SNP that is confirmed by SNP testing is usually designated in a green colored text or with a plus symbol ( + ) after the SNP designation. It is often referred to as derived or given a positive ( + ) code.
A tested SNP that is not confirmed or that is negative ( - ) is usually in red text.
SNPs pending testing, or assumed negative have different colors, but do not use the code of either positive ( +) or negative ( - ). Different colors can be used by various testing companies, so it is important to confirm the designations within the charts given.
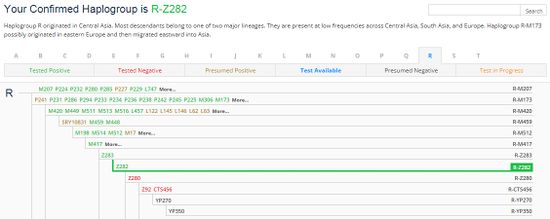
For an example of Haplogroup R1a1a1b1a with SNP marker of Z282 and using the shorthand code of R-Z282 uses designated colors representing the SNP status. See the image.
Use and importance of SNPs
Variations in the DNA sequences of humans can affect how humans develop diseases and respond to pathogens, chemicals, medication, vaccines, and other agents. SNPs are also thought to be key enablers in realizing the concept of personalized medicine. However, their greatest importance in biomedical research is for comparing regions of the genome between cohorts (such as matched cohorts with and without a disease).
Further reading
- The ABCs of DNA - SNPs by Kasandra Rose, DNA Footprints, 24 December 2016.
- DNA Heritage masterclass on SNPs and haplogroups (Internet Archive link)
Resources
- The ISOGG SNP index
- Human Genome Project Information — SNP Fact Sheet
- NCBI resources — Introduction to SNPs from NCBI
- The SNP Consortium LTD — SNP search
- SNPedia - a wiki devoted to the medical consequences of DNA variations, including software to analyze personal genomes
- HGNC Guidelines for Human Gene Nomenclature
- David Reynolds' SNP compendium
- Genetic variation databases - a resource from the Center for Human and Clinical Genetics with links to websites for: SNP databases, SNP-specific primer design software; SNP detection and effect prediction; and disease-causing variations.
Databases
- 1000 Genomes Project — A Deep Catalog of Human Genetic Variation
- NCBI dbSNP database — "A central repository for both single base nucleotide substitutions and short deletion and insertion polymorphisms"
- GWAS Central — GWAS Central (previously known as the Human Genome Variation database of Genotype-to-Phenotype information) is a central repository of findings from genome-wide association studies (GWAS). It includes information on all known SNPs and other variants from public databases such as dbSNP and the DBGV.
- PharmGKB — The Pharmacogenetics and Pharmacogenomics Knowledge Base, a resource for SNPs associated with drug response and disease outcomes
- International HapMap Project — "A public resource that will help researchers find genes associated with human disease and response to pharmaceuticals"; note that the HapMap Project was discontinued June 2016, but archived data is still available via FTP transfer from NCBI
Useful tools for advanced users
- SNPStats — SNPStats, a web tool for analysis of genetic association studies
- Restriction HomePage — a set of tools for DNA restriction and SNP detection, including design of mutagenic primers
- SIFT — "An online tool that predicts on the effect of SNPs on protein function"
- WatCut — an online tool for the design of SNP-RFLP assays
- Y-SNP converter