Cladogram
From ISOGG Wiki
In genetic genealogy, a cladogram is a diagram showing genetic relationships among a group of people. It depicts their commonalities and differences. "Phylogenetic tree" is a synonym for cladogram.
The chief value of cladograms is their graphic nature. A picture is often worth many words or numbers.
One example of a cladogram is the ISOGG phylogenetic tree of the human Y chromosome. Another is a descendancy chart printed by genealogical software.
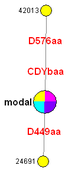
Another example is the Fluxus Network diagram to the right -- a cladogram depicting six persons' 37 Y-STR marker haplotypes for a genetic family in one surname DNA project. It consists of six taxa (plus the software-added "modal" shown in magenta; three have an exact match at 37 markers. There are two branches; one is two mutations from the modal and one is a different mutation from the modal.
Contents
Cladogram Basics
A cladogram applies cladistic techniques to genetic genealogy and pictorially displays the results. Cladistics "...is an approach to biological classification in which organisms are grouped together based on whether or not they have one or more shared unique characteristics that come from the group's last common ancestor and are not present in more distant ancestors. Therefore, members of the same group are thought to share a common history and are considered to be more closely related".[1]
Cladograms for genetic genealogy differ from those used in taxonomy in focusing only on humans and concerning smaller variations (e.g., specific STR or SNP mutations). Also, data for the root (ancestor) is typically not available and must be hypothesized.
In surname Y-DNA projects, cladograms can give an indication of which branches are likely to be more closely related to each other (and thus share a common ancestor more recently than other branches). This helps focus further traditional genealogical research in that it helps identify those branches which perhaps should be working more closely together to find documentary evidence of their (more recent) connection.
The basic unit of a cladogram is a taxon (plural taxa), an individual, group or species (in short, anything the cladisticist chooses). In Y-DNA cladograms, a taxon may be an individual's haplotype or pattern of STR markers and their allele values. The taxa in a cladogram are usually inferred to be phylogenetically related. It is recommended that the cladisticist assure phylogenetic relationships before constructing a cladogram.
A frequent cladogram convention is to imply "advancement" by direction. Upward and/or rightward may signify greater complexity; downward and/or leftward, less complexity. This may not apply in genetic genealogy; a mutated haplotype is not necessarily either "better" or "worse" than the prior form.
Cladograms often rely on a parsimony criterion (e.g., Occam's razor) to infer phylogeny from molecular data. That is, the explanation requiring the fewest assumptions is most likely to be true.
- It is assumed that haplotypes with greater similarity are more closely related than those with more dissimilarity.
- And in Y-STR terms, a group's modal marker value is the assumed value of the common ancestor, i.e., the "root".
Cladograms may have a wide range of complexity, depending on the number of taxa and the closeness of relationships. For example, an exact Y-DNA match between two or more people results in a single point (taxon). As differences increase, the points become separated.
Cladogram Benefits
Cladograms have uses in both analysis of genetic data and in illustrating the interpretation of the data.
- Analysis: Cladograms isolate certain aspects by ignoring properties shared in common and highlighting differences. Also, some types of cladogram may help discover selection bias in the data.
- Illustration: Pictures are simpler for many to understand than words or numbers. A cladogram may help to illustrate key points.
Cladogram Examples
Some cladogram examples:
Phylip
To the right is an example of Phylip output, in Newick tree format.
The (hypothesized) root is at the bottom left of the diagram.
Mutations are indicated by vertical lines; branches by horizontal lines.
Time scale is indicated below the tree.
Fluxus
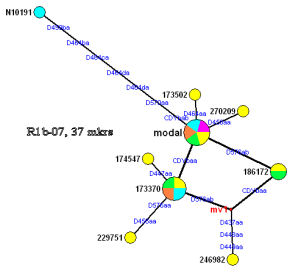
A more complex genetic family of 16 individuals; the "root" is at the node labeled "modal". Multiple exact matches in three nodes are represented by various-colored pie slices; by inference, these persons are more closely related to each other than to others in the group. Higher resolution (e.g., 67 markers) may show differences to separate the taxa into different nodes. The blue circle in the upper left corner represents a man whose MRCA with the others must predate the mid-1600s; here, genetic data and paper trails are consistent.
- The node labeled 173370 represents a major branch with two (or three) sub-branches.
- The node labeled "mv1" represents alternative descent paths -- from either 173370 or 186172. Additional testing may resolve the ambiguity.
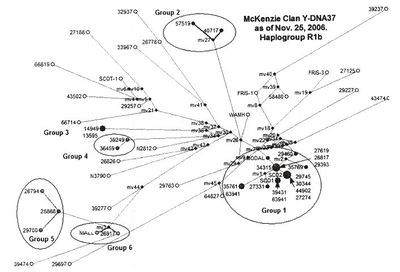 The Clan MacKenzie cladogram, from page 15 of http://www.electricscotland.com/mackenzie/images/news394.pdf displays additional complexity.
The Clan MacKenzie cladogram, from page 15 of http://www.electricscotland.com/mackenzie/images/news394.pdf displays additional complexity.
It was initially produced by Fluxus Network and subsequently modified. Note the circling of "Group 1", "Group 2", etc.
Hand-Drawn
It is possible to draw cladograms without the use of special software. (See the Robb article cited under "Further Reading".) Or, one may use drawing software. This approach may be necessary if one wants to combine genetic with genealogical data.
Cladogram Criticisms
Cladograms are not all-purpose tools; a cladogram may not show show what one expects.
Some specific criticisms:
- I didn't learn anything I didn't already know. Good; you've confirmed what you knew with an additional technique. Now you have a picture of it.
- The cladogram disagrees with the genealogical data. One should investigate which data is the more reliable and whether the genetic data has been input correctly.
- My exact matches are reduced to a single node (point, circle, block, etc.) This happens because the cladogram superimposes one taxon on anther. (A cladogram is a flat, two-dimensional representation of phenomena which, likely, exist in more dimensions than two.) Exact matches have no differences, so occupy the same two-dimensional space. (It is roughly analogous to a cladogram for primate species with two chimpanzees on the chart.) It may be possible to show the multiple taxa by color-coding "pie slices".
- My cladogram is too complicated; I can't make head nor tail of it. Perhaps, you've asked it to do too much. Try scaling back the objective and reducing the data set to fit.
- The software is poorly documented. This is a typical problem of freeware, particularly for specialized fields; developer altruism has a limit which often falls short of user manuals and help files.
- The underlying assumptions and processing are unstated. The publisher may be relying on your knowledge of computational cladistics to fill in the gaps. Many users are doctoral students and expert researchers in this field, for whom explanation is unneeded.
Constructing a Cladogram
The steps in constructing a cladogram are:
- Gather and organize the data. This will often be Y-STR allele values by marker. The Y-STR must all be tested at the same resolution.
- Consider possible cladograms. This may involve specialized software such as Fluxus or PHYLIP. McGee Utilities can be used to format your STR data for input. (Copy and paste the McGee output into a plain-text file.)
- Fluxus may be downloaded free. An acknowledgement is required when publishing diagrams or output data. The illustration above shows a simple version of its output.
- PHYLIP is a set of 35 programs, also free but designed for academic use. Output from "drawtree" is in the "Newick" tree format.
- Other cladogram software is listed at http://evolution.genetics.washington.edu/phylip/software.pars.html.
- Select best cladogram - the one that most closely portrays the data and illuminates the relationships.
Further reading
- Fluxus software
- Phylip software
- Dave Hamm's instructions on making cladograms
- Fluxus network diagrams vs hand-constructed Mutation History Trees by John Barrett Robb, www.johnbrobb.com, 15 April 2012.
- Identifying sub-groups in the Borders Family: TIPs, SNPs and cladograms A paper by James M Irvine discussing the use of Fluxus software in the Irwin DNA Project, published April 2011, revised June 2011.
- Cladistics - revealing the tree of life
Acknowledgements
Thanks to ISOGG member Ralph Taylor for providing the text for this page.
See also
References
- ↑ Wikipedia article on cladistics. Accessed 12 December 2013.