Fully identical region
From ISOGG Wiki
Two individuals are said to share a fully identical region (FIR) or completely identical region of DNA if the alleles on both chromosomes are identical throughout the region, in other words if either (a) one individual's paternal chromosome is identical to the other individual's paternal chromosome and their maternal chromosomes are also identical throughout the region or (b) one individual's paternal chromosome is identical to the other individual's maternal chromosome throughout the region and vice versa.
In practice, it is not always possible to distinguish whether the relationship is fully identical by descent or fully identical by chance. For example, if the SNPs in John's paternal and maternal chromosomes are AAAAAAAAAA and GGGGGGGGGG respectively and the SNPs in Mary's paternal and maternal chromosomes are AGAGAGAGAG and GAGAGAGAGA respectively, then ten consecutive AGs will be observed for both individuals and John and Mary will appear to be fully identical in this region when in fact they are not when the data is not phased.
Identical twins are fully identical at every point in their DNA. Other full siblings, including non-identical twins, share around 50% of their DNA, and have both half-identical regions and completely identical regions. The expected numbers for full siblings are 50% half-identical, 25% completely identical, and 25% not identical for an overall average of 50%. Half-siblings do not in general share any fully identical regions. Double cousins who are related through both their paternal and maternal lines also share some fully identical regions. It is also possible that small FIRs will be shared by more distantly related individuals whose parents are all from the same highly endogamous (inbred) population.
Neither 23andMe nor Family Tree DNA distinguishes between fully identical regions and half-identical regions when computing the number of shared centiMorgans (cMs). Thus, for example, the shared cM for full-siblings will on average be 75% of the total length of the genome, of which on average 50 percentage points are half-identical and 25 percentage points are fully identical.
Similarly, the FTDNA Family Finder chromosome browser and the chromosome browser in 23andMe's Family Inheritance: Advanced feature do not distinguish between fully identical and half-identical regions. However, it is possible to see the fully identical regions at 23andMe by using the Family Traits chromosome browser (accessed via the Family and Friends menu). AncestryDNA does not provide a chromosome browser or information on matching segments. Those who have tested at Family Tree DNA or AncestryDNA can see their FIRs only by uploading their raw data to the free GedMatch utility. Fully identical regions show up in green in the one-to-one comparison, though in the one-to-many results at GEDMatch no distinction is drawn between FIRs and HIRs.
There are usually no start or end points listed for the fully identical regions between siblings. These position numbers can be very important in phasing. Some of this information can be obtained by uploading the raw data to David Pike's utilities. This is not a browser and gives information in megabases instead of centiMorgans but it was the first useful tool available for making length comparisons in fully identical regions.
Contents
Chromosome browser examples of fully identical regions
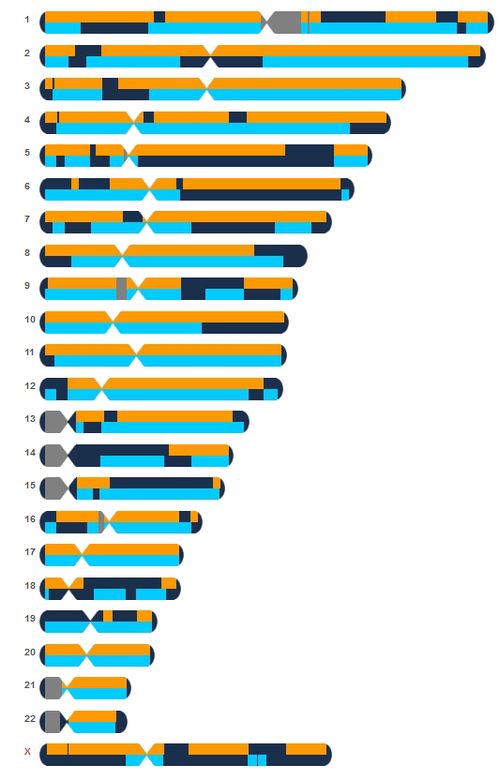
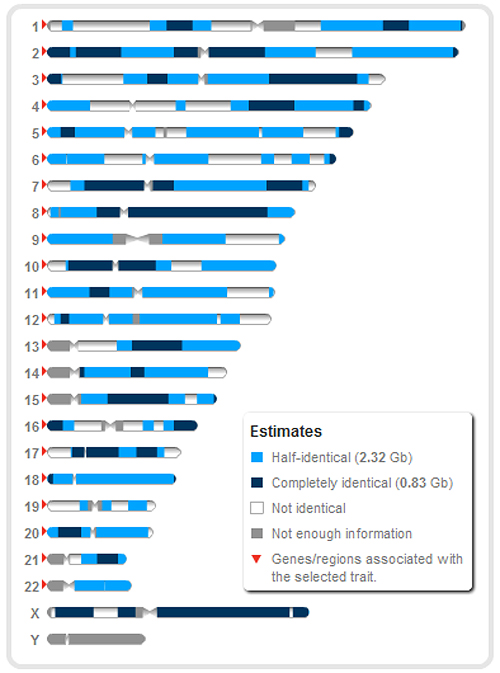
The following screenshot shows a comparison between two brothers at 23andMe. The fully identical regions are shown in dark blue, the half-identical regions are shown in light blue and the non-identical regions are shown in white:
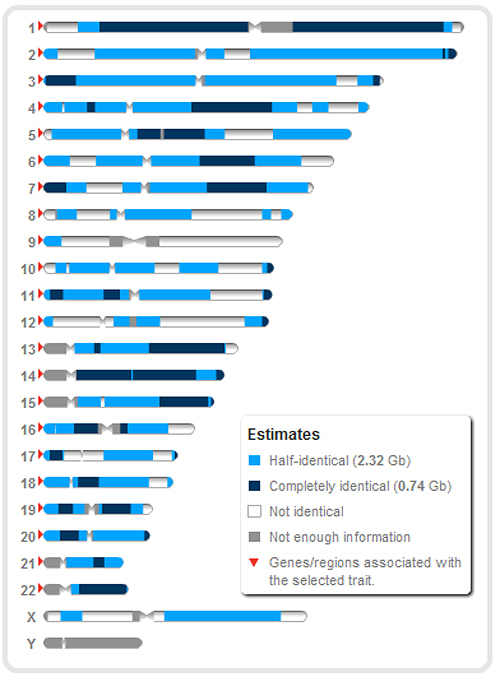
The following is an example of the sharing between a brother and sister at 23andMe:
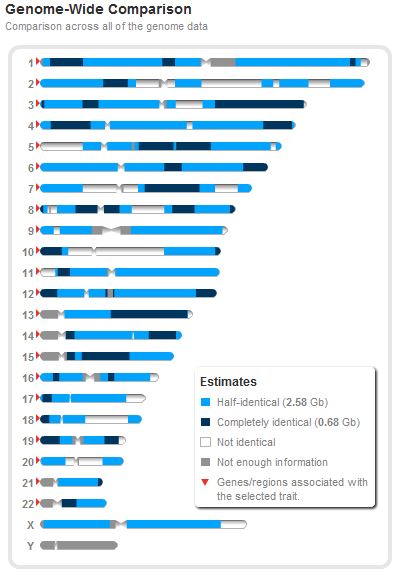
The following is another example of the sharing between a brother and sister at 23andMe. They share 54.4% of their DNA (3154 cMs, 39 segments):
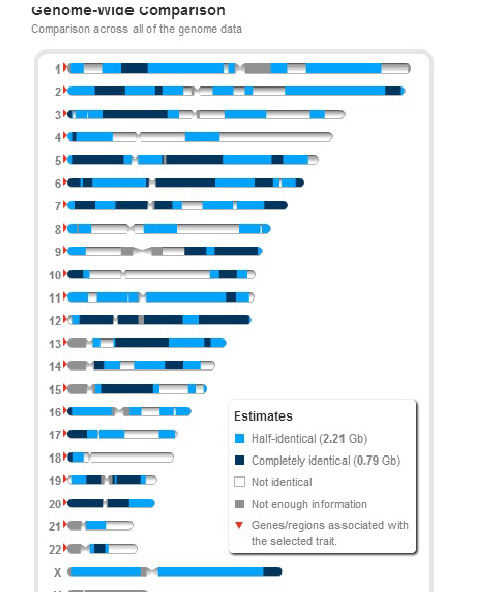
The following is an example of the sharing between two sisters at 23andMe:
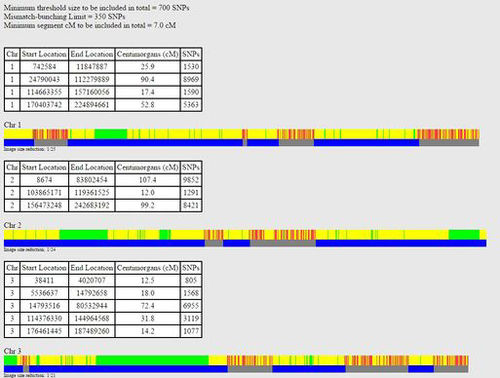
The following is an excerpt from a one-to-one comparison between two siblings at GedMatch. The fully identical regions are shown in green, half-identical regions are shown in yellow, and non-matching regions are shown in red. The large blue blocks indicate matching segments of 7 cMs or more.
The following is an excerpt from a one-to-one comparison between two double cousins at GedMatch. They are paternal first cousins and maternal second cousins. The fully identical regions are shown in green, half-identical regions are shown in yellow, and non-matching regions are shown in red. The large blue blocks indicate matching segments of 7 cMs or more.
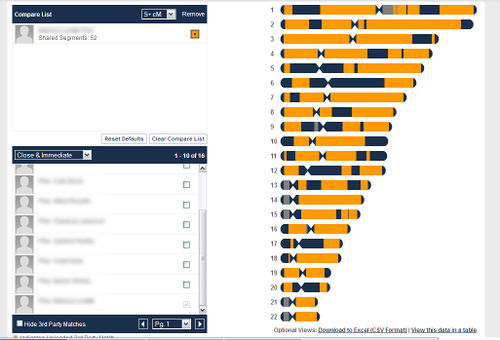
The following is a comparison between two siblings who have taken the Family Finder test at Family Tree DNA. Note that the browser does not distinguish between half-identical regions and fully identical regions.
The following is a comparison between an individual who has taken the Family Finder test at Family Tree DNA and two of her full siblings.
Further reading
- Relatedness A helpful article in the Understanding Genetics series provided by Stanford at the Tech. The article explains the basics of recombination and inheritance, and also includes chromosome browser diagrams from 23andMe showing half-identical and fully identical regions. (See Girl vs. Brother 1 and Girl vs. Brother 2 for examples of fully identical regions.)
- Family Traits: Genome View An article from 23andMe with explanations of the inheritance of DNA segments. Chromosome browser examples are provided for siblings, grandparent versus grandchild and parent vs child.
- Can 23andMe distinguish half vs full sibling relationships?
Blog posts
- Obtaining FIR boundaries on GEDmatch using phased and "My Evil Twin" phased kits by Sue Griffith. Genealogy Junkie,4 December 2016.
- Obtaining FIR boundaries on GEDmatch using the little tick marks by Sue Griffith. Genealogy Junkie, 1 November 2016.
- Full versus half sibling DNA matches by Kitty Cooper, Kitty Cooper's Blog 29 April 2016.
- Questions from my inbox: siblings by Andrea Badger, Andrea's Ancestors, 8 December 2011.