Cousin statistics
From ISOGG Wiki
It is helpful to have an understanding of the probabilities of matching cousins at different degrees of relationship when contemplating autosomal DNA testing or when interpreting the results of an existing test. This article reports some relevant cousin statistics.
Company statistics
The following table shows the information provided by the three main testing companies on the probability that two cousins will share enough DNA for the relationship to be detected. AncestryDNA claim to be able to detect more relatives in the third to sixth cousin range which possibly relates to their use of phasing. Note that each company sets its own autosomal DNA match thresholds and some more distant matches might not appear because of these restrictions.
| Relationship | 23andMe[1] | AncestryDNA[2] | Family Tree DNA Family Finder[3] |
|---|---|---|---|
| First cousins | 100% | 100% | 100% |
| Second cousins | 100% | 100% | >99% |
| Third cousins | 89.7% | 98% | >90% |
| Fourth cousins | 45.9% | 71% | >50% |
| Fifth cousins | 14.9% | 32% | >10% |
| Sixth cousins | 4.1% | 11% | Remote (typically less than 2%)[4] |
| Seventh cousins | 1.1 | 3.2% | |
| Eighth cousins | 0.24 | 0.91% | |
| Ninth cousins | 0.06% | ||
| Tenth cousins | 0.002% |
Theoretical probabilities
Amy Williams and colleagues at Cornell University have provided two useful chart based on simulations which shows the likelihood of matching with cousins of different degrees of relationship and the number of segments shared:
- How often to do two relatives share DNA? 3 November 2020
- How often do two half-relatives share DNA/ 6 November 2020
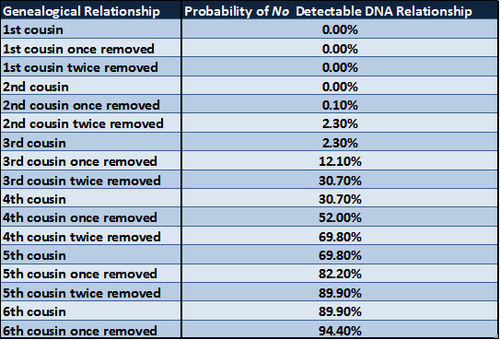
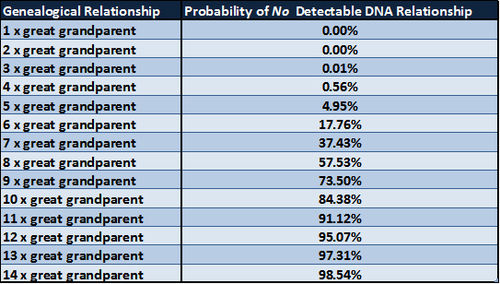
Simulations in a 1983 paper by Kevin Donnelly showed the theoretical probability of having no detectable DNA inherited from a specific ancestor and the probability of sharing DNA with cousins of varying degrees of relationship. The content of the following two tables is derived from Table 1 in the paper The probability that related individuals share some section of genome identical by descent by Kevin P Donnelly, Statistical Laboratory, Cambridge University, Cambridge, England. (Source: Theoretical Population Biology 1983: 23, 34-63) A copy of the paper is available here.
In their research presented in the February 2016 issue of Theoretical Population Biology[5], M. Sun, N.A. Sheehan, et al. conducted extensive simulations based on actual test results taken from modern microarray autosomal DNA tests and taking into account the effect of linkage disequilibrium. The authors examined the probability that any two people, via SNP testing, could be detected to have shared genetic ancestry versus the likelihood that they might be completely unrelated. For specificity, the models were run assuming there was a single MRCA, not a couple; i.e., that the ancestral lines descended from half-sibling children of the MRCA.
The authors organized the runs using four different densities of SNPs tested, with 500K being the closest value to our common microarray tests. In the table below the probability averages have been converted to percentages, and it is important to note that a value of 50% indicates at that level the ability to predict shared genetic ancestry as opposed to unrelatedness is indistinguishable per the simulation parameters:
| No. of SNPs | Siblings | 1st Cousins | 2nd Cousins | 3rd Cousins | 4th Cousins | 5th Cousins |
|---|---|---|---|---|---|---|
| 2,200 | 100% | 92.5% | 60.5% | 51.5% | 50% | 50% |
| 22,000 | 100% | 100% | 94.7% | 68.5% | 55% | 54.7% |
| 500K | 100% | 100% | 100% | 87.8% | 61.2% | 55.1% |
| 1 million | 100% | 100% | 100% | 87.2% | 64.7% | 55.7% |
One conclusion of the authors was, "as expected, identification of the true relationship from unrelated becomes more difficult as the relationship becomes more distant. We also confirmed...that it is much more challenging to distinguish between the true pedigree and a close alternative structure (rather than just 'unrelated') after about 6 separating meioses [2nd cousins] with 500K SNPs."[5] See also Table 3, Skare, Sheehan, and Egeland (2009)].[6]
How many cousins do we have?
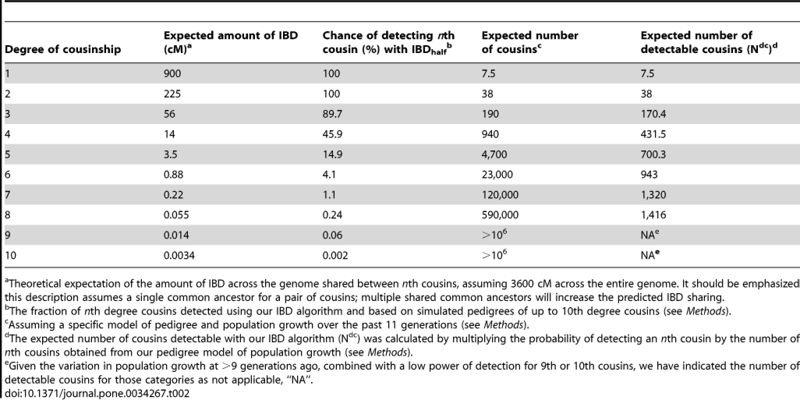
Although there is only a low chance of sharing enough DNA with a specific distant cousin for the relationship to be detected, we have a large number of distant cousins and so many of these more distant cousins will appear in our match lists. The following table from the paper Cryptic distant relatives are common in both isolated and cosmopolitan genetic samples by Henn et al (2012) shows the expected number of cousins at different degrees of relationship and the expected number of detectable cousins along with the expected amount of Identical by descent (IBD) sharing if the relationship is detected.
Mathematician and genetic genealogist Paul Rakow has done his own computer simulations on family sizes and has published the results in an essay on Counting cousins (published online 31 March 2016).
A study by AncestryDNA, based on British birth rates, census data, parliamentary research briefings[7] and other sources for the last 200 years, produced the following statistics on the number of cousins that the average British person would be expected to have.[8][9]
| Relationship | Number of cousins |
|---|---|
| First cousins | 5 |
| Second cousins | 28 |
| Third cousins | 175 |
| Fourth cousins | 1,570 |
| Fifth cousins | 17,300 |
| Sixth cousins | 174,000 |
It is not clear if these statistics relate to the whole of the United Kingdom or just England and Wales.
Further reading
- How often to do two relatives share DNA? by Amy Williams HAPI-DNA, 3 November 2020.
- How often do two half-relatives share DNA by Amy Williams HAPI-DNA, 6 November 2020
- IBD (identical by descent sharing rates by cM length by Amy Williams HAPI-DNA, 12 November 2021
- Why your genetic tree is not the same as your family tree by Diahan Southard, Lisa Louise Cooke's Genealogy Gems, 7 May 2017.
- What are you chances of finding an autosomal DNA match? by John Reid, Canada's Anglo-Celtic Connections, 3 October 2016.
- How many cousins by John Reid, Canada's Anglo-Celtic Connections, 14 August 2016.
- How many cousins share my 5th grandparents? by Kitty Cooper, Kitty Cooper's Blog, 19 July 2016.
- Face it: DNA cannot find all your relatives. DNA.Land blog, 27 February 2016.
- Your family: past, present and future by Tim Urban, Wait But Why.
- How many genetic ancestors do I have? by Graham Coop, The Coop Lab blog, 11 November 2013.
- Widen the net by Judy Russell, The Legal Genealogist, 7 April 2013. A cautionary tale about third cousin matches.
- Q&A: Everyone has two family trees - a genealogical tree and a genetic tree by Blaine Bettinger, The Genetic Genealogist, 10 November 2009.
Tools
- The Shared cM Project 4.0 tool v4 Interactive version by Jonny Perl
See also
- Autosomal DNA portal
- Autosomal DNA match thresholds
- Autosomal DNA statistics
- Coefficient of relationship
- Cousin
- Pedigree collapse
Acknowledgements
Thanks to Jon Hamm for compiling the spreadsheets based on data from Donnelly (1983) that have been used in the section on theoretical probabilities.
Footnotes
- ↑ The 23andMe data were extracted from the paper by Henn et al. "Cryptic distant relatives are common in both isolated and cosmopolitan genetic samples" published online in PLOS One on 3 April 2012. 23andMe has a simplified version of these data in a customer FAQ "The probability of detecting different types of cousins".
- ↑ The AncestryDNA data were extracted from information provided in the first "AncestryDNA Matching White Paper," 2014, in section 5.3 "Relationship Estimation Evaluation." These calculations used small segment sizes but as of 31 August 2020 AncestryDNA has stopped reporting on matches sharing less than 8 cM total.
- ↑ The Family Finder percentages are from an article in the Family Tree DNA Learning Center "What is the probability that my relative and I share enough DNA to be detected by Family Finder?"
- ↑ The figure quoted by Family Tree DNA refers to "6th cousins and more distant".
- ↑ 5.0 5.1 M. Sun, N.A. Sheehan, et al., "On the Use of Dense SNP Marker Data for the Identification of Distant Relative Pairs." Theoretical Population Biology 107 (February 2016): 14-25. DOI: https://doi.org/10.1016/j.tpb.2015.10.002
- ↑ Øivind Skare, Nuala Sheehan, Thore Egeland, "Identification of Distant Family Relationships." Bioinformatics 25:18 (September 2009) 2376–2382. DOI: https://doi.org/10.1093/bioinformatics/btp418
- ↑ A Century of Change: Trends in UK statistics since 1900 House of Commons Research Paper 99/111. Published 21 December 1999.
- ↑ Average British person has 193,000 living cousins says new research. Daily Mirror. 17 June 2015.
- ↑ Ancestry. There's a one in 300 chance that a complete stranger is your cousin. Ancestry blog, 26 June 2015. Also available as a press release on the Ancestry Corporate website: One in 300 chance that a complete stranger is your cousin.